We conduct discovery and applied research to improve the resistance of bread wheat against major diseases.

Wild wheat disease resistance genes
Collaborator: Open Wild Wheat Consortium
The development of agriculture ~10,000 years ago in the Fertile Crescent in the Middle East allowed the establishment of large, settled societies. Several millennia of cultivation and selection of the best-performing variants led to a suite of cereal crops and pulses better adapted to human requirements in terms of cultivation, yield, storage and taste. However, this domestication also introduced an Achilles heel by severely eroding genetic diversity. One of the most extreme bottlenecks occurred when cultivated tetraploid wheat (Triticum turgidum) hybridised with the wild goatgrass Aegilops tauschii to give rise to hexaploid bread wheat (T. aestivum).
The El Dorado of genetic diversity still present in Ae. tauschii represents a fascinating resource for fundamental studies in cereal biology and practical exploitation. In our recent work conducted under the aegis of the Open Wild Wheat Consortium, we used association genetics to pinpoint candidate genes for wheat performance and environmental resilience. We also characterized a rare, evolutionary distinct lineage of Ae. tauschii and demonstrated its previously unknown contribution to the bread wheat genome (Gaurav et al., 2022).
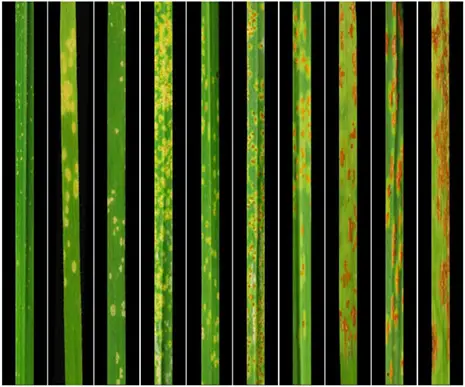
Figure 1. Phenotypic diversity in Ae. tauschii in response to Puccinia graminis f. sp. tritici, the causal agent of wheat stem rust (Arora et al., 2019)
Presently, we are constructing new genomics resources in Ae. tauschii that will permit (i) a global overview of genetic variants present across the species (including short sequence polymorphisms, structural alterations, epialleles and rare alleles) and (ii) accelerate the functional validation of candidate genes. This will allow us to address fundamental questions concerning the species-wide genetic heterogeneity that provides stress and disease resilience over evolutionary timescales. We are focusing on two traits that exhibit pronounced diversity in Ae. tauschii including (i) resistance to stem rust, an important disease of wheat (Figure 1); and (ii) trichomes, small leaf appendages involved in biotic and abiotic stress tolerance (Figure 2).
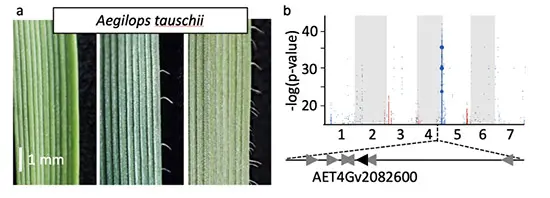
Figure 2. Phenotyping and genetic diversity for trichome formation in Ae. tauschii. a, Phenotypic variation for trichome number on leaves of 3-week-old Ae. tauschii plants. b, Genetic identification of a candidate gene controlling trichome density by k-mer-based association mapping (Gaurav et al., 2022)
We have also sequence-configured a genetically diverse panel of Ae. sharonensis, another species of goatgrass that diverged from Ae. tauschii ~2 million years ago. We are conducting comparative genomics analyses to shed light on the dynamics that shaped the functional resistance gene repertoire in the two species.

Mechanism of atypical wheat resistance genes
Collaborators: Simon Krattinger at KAUST, Łukasz Jaremko at KAUST, Ikram Blilou at KAUST, Heribert Hirt at KAUST
In plants, major dominant race specific resistance genes that have been defined by genetics and subsequently cloned often encode intracellular receptors of the nucleotide-binding and leucine-rich repeat class or extracellular immune receptor-like proteins. The guard hypothesis, a mechanistic framework first proposed in 1998 and since validated experimentally for several resistance proteins, dictates that the resistance protein guards the target of a pathogen virulence (effector) protein. Perturbation of this target by the effector is detected by the resistance protein leading to a conformational change that triggers downstream defence responses.
In recent years, a class of disease resistance genes encoding protein kinase fusion proteins has been identified in the Triticeae, the group of small grain cereals that include the major crops wheat, barley and rye. Members of this class of genes include Rpg1 from barley, Yr15H from wild emmer wheat, Sr60 from Einkorn wheat, and Pm4 from Persian wheat. We have added to this class with the cloning and characterization of WTK4 from Ae. tauschii, Rwt4 from bread wheat, Sr62 from Ae. sharonensis (Figure 1), and Sr43 from Thinopyrum ponticum. We are interested in elucidating the mechanism of Sr43 and Sr62 using molecular, biochemical and structural approaches.
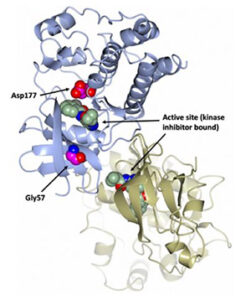
Figure 1. Homology model of Sr62. The overall structure of the tandem kinase is shown in ribbon representation with the N-terminal kinase domain in blue and the C-terminal kinase domain in gold (Yu et al., 2022).

Wheat pathogen genomics
To understand the evolution of wheat stem rust resistance genes and decipher the resistance mechanism in the host, we also need to identify the molecules (effectors) in the fungus that are recognized by the corresponding resistance proteins. We are using a mutational genomics approach to identify effectors corresponding to some of our key cloned resistance genes. We have identified EMS-induced mutants of P. graminis f. sp. tritici that have gained virulence on Sr43 or Sr45 plants (Figure 1). Cloned effectors can also be used as molecular tools to confirm the function of individual resistance genes that are part of polygene stacks. In this context, pathogen assays are not always informative due to the epistasis caused by the simultaneous interaction of multiple resistance genes with their corresponding effectors.
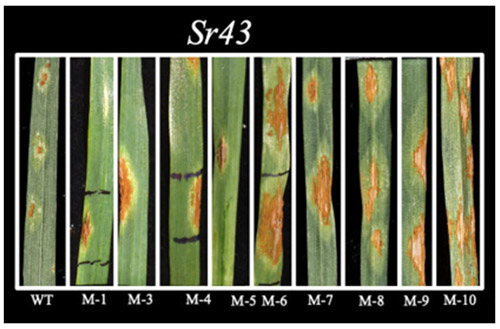
Figure 1. Nine EMS-induced mutants of P. graminis f. sp. tritici isolate UK-01, which have gained virulence on Sr43 plants (Kangara et al., 2020).

Engineering disease-resistant wheat
Collaborators: Wendy Harwood, Mark Smedley Sadiye Hayta at the John Innes Centre
Breeding for durable rust resistance can be achieved by combining multiple resistance genes into the same variety, a challenging exercise with conventional breeding, but now a viable proposition using ‘gene cassettes’ (Figure 1). In this approach, multiple resistance genes are assembled at the same location in the genome using biotech intervention, as recently demonstrated for five stem rust resistance genes in wheat (Luo et al., 2021).
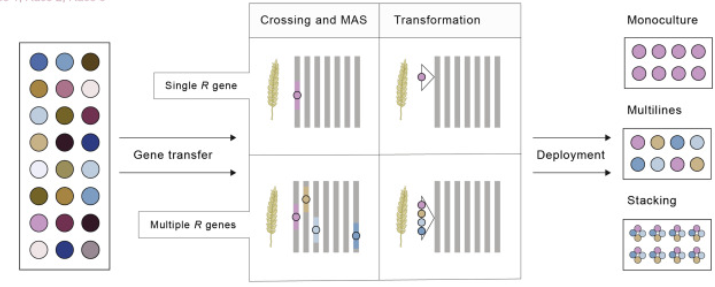
Figure 1. Resistance gene deployment using conventional and biotech breeding approaches (Hafeez et al., 2021).
Although >30 rust resistance genes have been cloned to date, providing the raw material necessary for engineering large cassettes, we lack a comprehensive picture of the efficacy of individual resistance genes against individual pathogen isolates representative of the genetic diversity in the pathogen population. This hinders the rational design of cassettes with multiple layers of resistance effective against all isolates of the pathogen, a prerequisite for durability.
Ascertaining the full resistance spectrum of cloned resistance genes is currently complicated by the multitude of different genetic backgrounds in which other resistance genes can mask the effect of the individual resistance gene under study. To address this knowledge gap, we plan to establish a Wheat Resistance Gene Library. This will include (i) the synthesis of cloned resistance genes, (ii) the transformation of these genes individually into a universally susceptible wheat cultivar for the respective rust pathogen, (iii) selection of genetically stable lines, (iv) deposition of the constructs and germplasm in public repositories, and (v) wide-scale phenotyping of the isogenic resistance gene lines.
